SPACEc
A Stanford academic library for multiplexed imaging analysis — from cell segmentation to spatial analysis.
19,123 lines of Python · 6 core modules · 19 notebooks · Published on PyPI & bioRxiv

A streamlined, interactive Python workflow for multiplexed image processing — from tissue extraction to spatial analysis in a single library.
sp.tl.cellpose_segmentation(img, ...)
sp.pp.filter_cells(adata, ...)
sp.tl.leiden(adata, resolution=1.0)
sp.tl.annotate(adata, method='stellar')
sp.tl.patch_proximity(adata, ...)
sp.pl.spatial(adata, color='cell_type')
The Problem
Multiplexed imaging lets researchers see 40+ proteins in a single tissue section — a massive leap for understanding how cells interact in cancer, autoimmunity, and neuroscience. But the analysis pipeline is duct tape: Cellpose for segmentation, scanpy for clustering, custom scripts for spatial analysis, matplotlib for visualization.
Researchers were spending more time wrangling code than doing science. Each lab had its own fragile collection of notebooks that broke when dependencies updated.
The Idea
A unified library that handles the entire workflow — raw TIFF images to spatial analysis results — in a single, consistent API. SPACEc follows the scanpy convention: sp.tl (tools), sp.pp (preprocessing), sp.pl (plotting), sp.hf (helpers). Built on AnnData objects, the standard format in single-cell biology.
Architecture
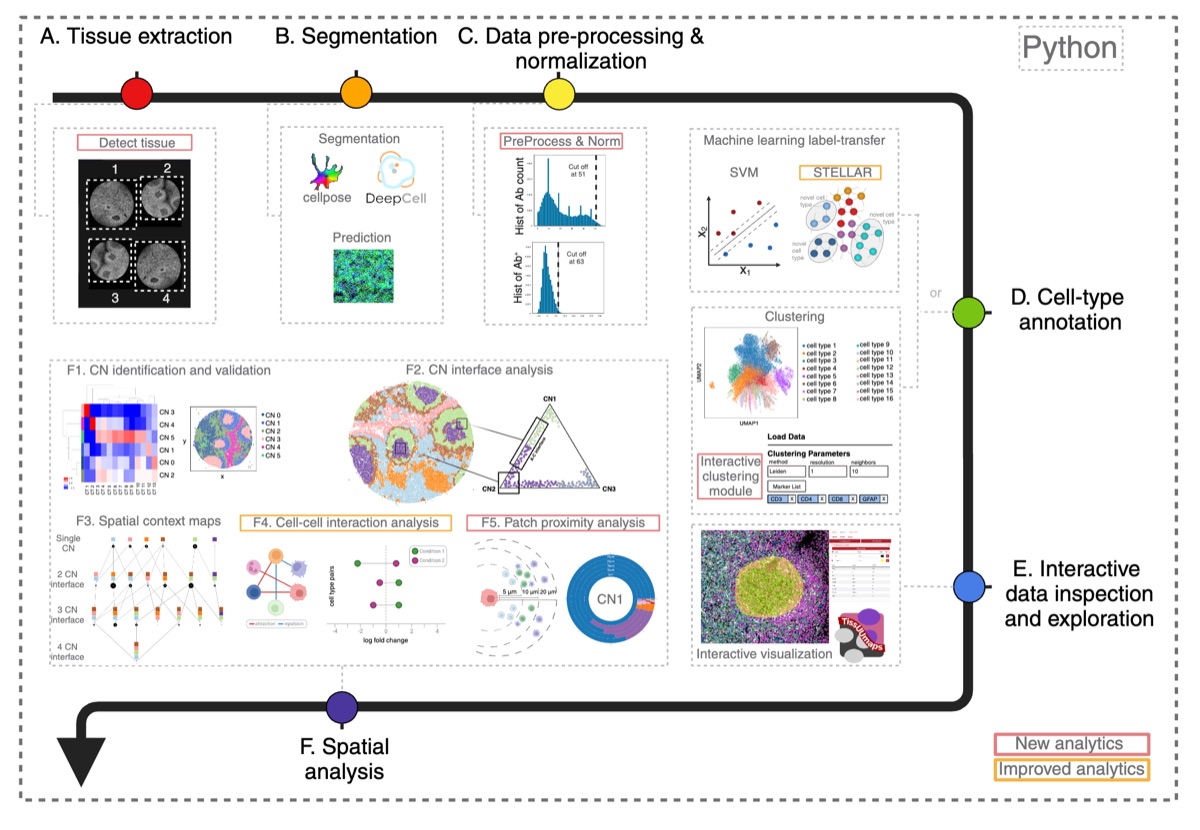
Full pipeline overview from the SPACEc paper
TIFF images
CSV markers
DeepCell (Mesmer)
QuPath import
Noise removal
Normalization
FlowSOM
GPU (RAPIDS)
STELLAR (GNN)
Hyperparameter tuning
Cell interactions
Patch Proximity*
*Patch Proximity Analysis is the novel research contribution.
Module Structure
- sp.tl._general 5,446 lines — core analysis, clustering, annotation, spatial
- sp.pl._general 5,214 lines — all visualization and plotting
- sp.tl._segmentation 2,522 lines — Cellpose, DeepCell, image processing
- sp.hf._general 1,096 lines — utilities and helper functions
- sp.pp._general 762 lines — preprocessing pipelines
- sp._shared.segmentation — shared segmentation logic
The Novel Contribution
Patch Proximity Analysis
Traditional neighborhood analysis looks at individual cell-to-cell distances. It tells you that cell A is near cell B, but it misses the bigger picture: how are groups of cells organized at the tissue level?
Patch Proximity Analysis works differently. First, DBSCAN identifies spatial clusters of a given cell type — patches of tumor cells, immune cell aggregates, stromal clusters. Then concave hull algorithms draw tight boundaries around each patch. Finally, the analysis measures how other cell types relate to those boundaries: which cells are inside? Which are at the border? Which are nearby?
This captures tissue-level spatial organization that cell-level analysis misses entirely. A tumor microenvironment isn't just individual interactions — it's the architecture of which cell populations are adjacent to which other populations.
Decisions
Why AnnData?
AnnData is the standard data structure in single-cell biology. By building on it, SPACEc integrates immediately with scanpy, the dominant analysis framework. Researchers don't learn a new format — they move between SPACEc and scanpy seamlessly. The .obs, .var, .obsm structure maps naturally to cell metadata, marker metadata, and spatial coordinates.
Why support both Cellpose and DeepCell?
Different imaging modalities need different segmentation approaches. Cellpose excels at cytoplasm segmentation with limited training data. DeepCell (Mesmer) is better for nuclear segmentation in whole-cell multiplex images. Supporting both means researchers pick the best tool for their data without leaving the library.
Why GPU acceleration?
Multiplexed imaging datasets are enormous — tens of thousands of cells per sample, hundreds of samples per study. CPU-based Leiden clustering that takes 20 minutes drops to under 2 minutes on GPU via RAPIDS. For iterative analysis, that's the difference between interactive exploration and going for coffee.
Why the scanpy API convention?
Familiarity reduces adoption friction. Every computational biologist already knows sc.tl, sc.pp, sc.pl. Using the same pattern (sp.tl, sp.pp, sp.pl) means the learning curve is the domain, not the API. It's a small decision that makes a large difference in whether people actually use the library.
Tradeoffs
Python 3.9–3.10 only. TensorFlow, PyTorch, and RAPIDS all have strict version requirements that don't overlap cleanly above 3.10. This is the single biggest friction point for adoption. Docker containers (CPU and GPU variants) are the recommended install path for a reason.
Monolithic modules. _general.py at 5,446 lines is hard to navigate and contributes to. The scanpy convention prioritizes API consistency (sp.tl.function()) over file organization. This was a conscious tradeoff — the public API is clean even if the internals are dense.
Dependency sprawl. Cellpose, DeepCell, RAPIDS, PyTorch Geometric, GeoPandas, TensorFlow — each brings transitive dependencies that conflict with each other. A fresh install without Docker is an afternoon project. This is the cost of wrapping the best tool for each step rather than reimplementing everything.
Talk
